Introduction
Whole genome sequencing is a method to analyze a specific polymorphism from a species with a known reference genome. The method can discover molecular sarcoma and various genetic polymorphisms (SNPs, insertions, deletions, inversions, complex rearrangements, copy number variation) related to performance and other useful applications. For whole-genome sequencing, the combination of short inserts and longer reads allows the characterization of any genome. For de novo whole-genome sequencing, the unparalleled raw read accuracy of Illumina next-generation sequencing (NGS) technology provides high quality, long contig assemblies.
As a professional sequencing provider, Quick Biology performs whole genome sequencing on HiSeq X Ten (HiSeq X 10), which provides the most economical way for whole genome sequencing.
a Whole Genome Sequencing Quote, Click Here!
Bioinformatics for Whole Genome Sequencing:
a.Germline variants, compared to reference genome
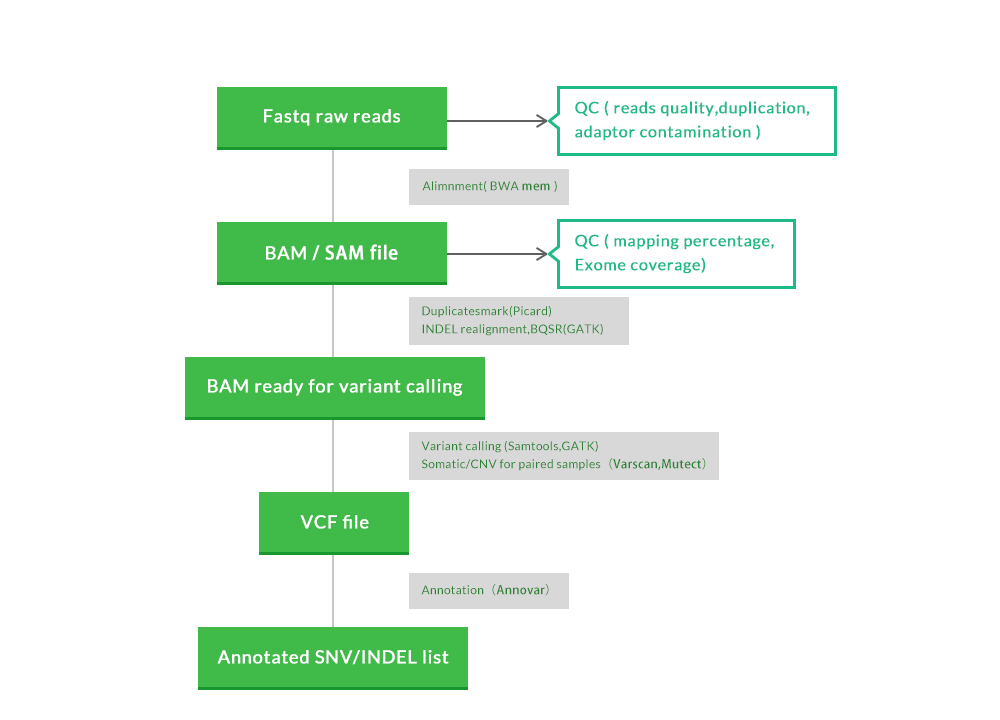
Workflow
Sample Requirment
DNA amount quantified by Nanodrop: ≥500 ng for normal input; minimum 20 ng
DNA concentration: ≥2 ng/μL
Purity: A260/280:1.8-2.0, A260/230: ≥ 1.8, Intact DNA, RNA-free, and non-degraded.
Turnaround Time
About 3-4 weeks, depends on the quality of submitted samples.